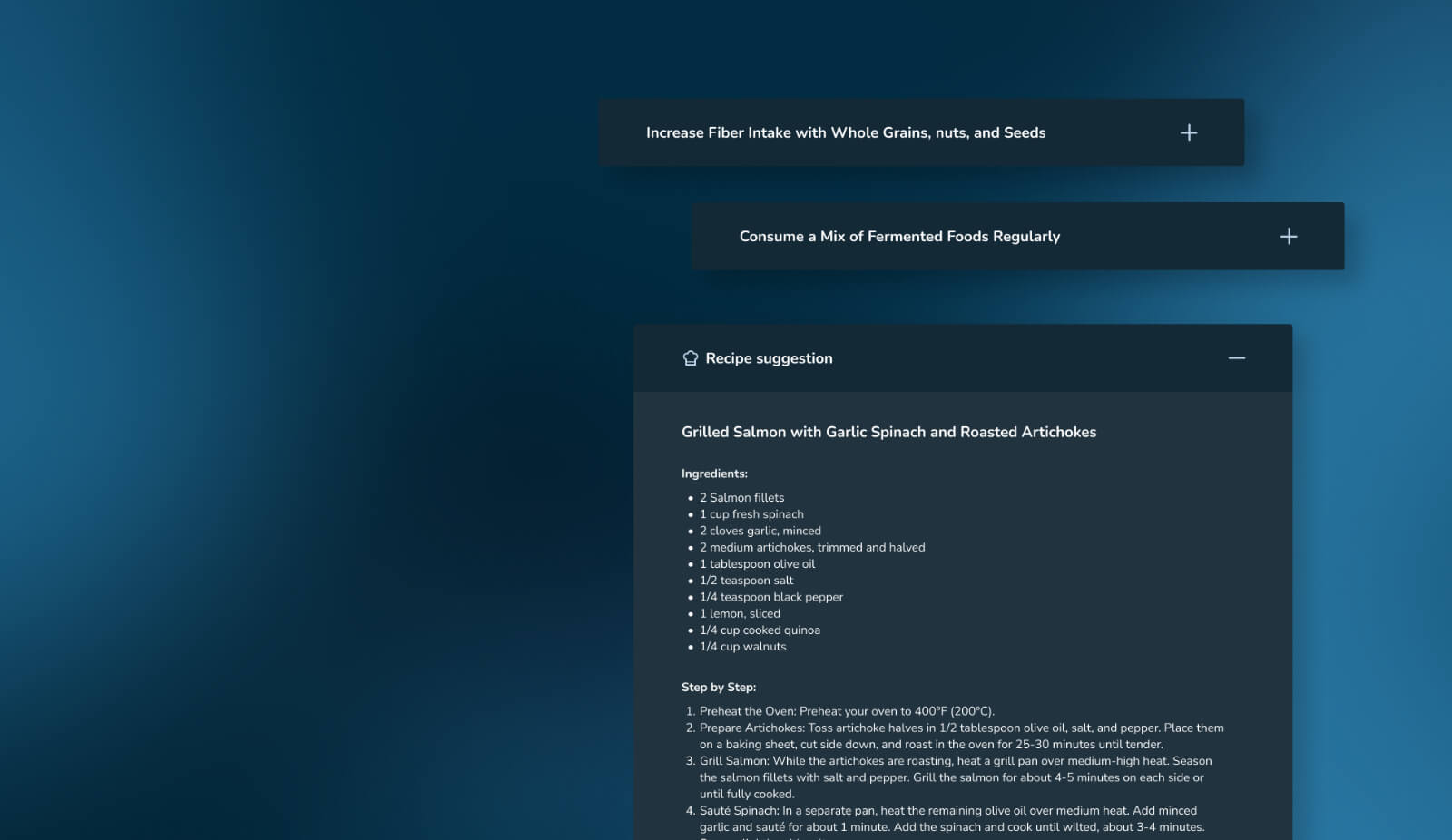
BiomeDx integrates seamlessly into laboratory workflows through multiple connection methods: API, sFTP, Illumina BaseSpace and other sequencer connectors.
This ensures compatibility with 99% of existing laboratory systems, enabling instant onboarding without technical overhead.
Science you can trust.
BiomeDx’s technology is backed by extensive research and clinical validation. Our findings have been published in 13+ peer-reviewed papers across leading journals including Nature Portfolio, Gut Microbes, and Frontiers in Microbiology.
We are also the only microbiome diagnostics company globally certified to ISO 9001 & ISO 13485, ensuring medical-grade quality standards.
We are also the only microbiome diagnostics company globally certified to ISO 9001 & ISO 13485, ensuring medical-grade quality standards.



Assessing the Performance of a Novel Stool-Based Microbiome Test That Predicts Response to First Line Immune Checkpoint Inhibitors in Multiple Cancer Types
CANCERS
Using fecal immmunochemical cartridges for gut microbiome analysis within a colorectal cancer screening program
GUT MICROBES
Characteristics of Gut Microbiota in Rosacea Patients-A Cross-Sectional, Controlled Pilot Study
Life
Explore our research
Visit our newsroom